FULL TITLE:
Detailed Mapping of Human Habenula Resting-State Functional Connectivity
SPECIES:
Human
DESCRIPTION:
These scenes correspond to the main and supplementary results figures from our study of human habenula (Hb) resting-state functional connectivity using data from the 3T HCP young adult cohort. Scripts to generate subject-level fMRI ROIs optimized for Hb BOLD sensitivity are available at https://github.com/junqianxulab/Habenula_fMRI_ROIs.
ABSTRACT:
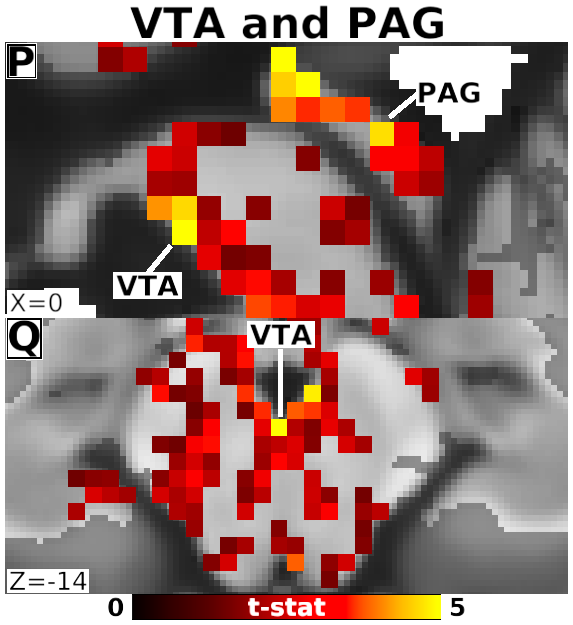
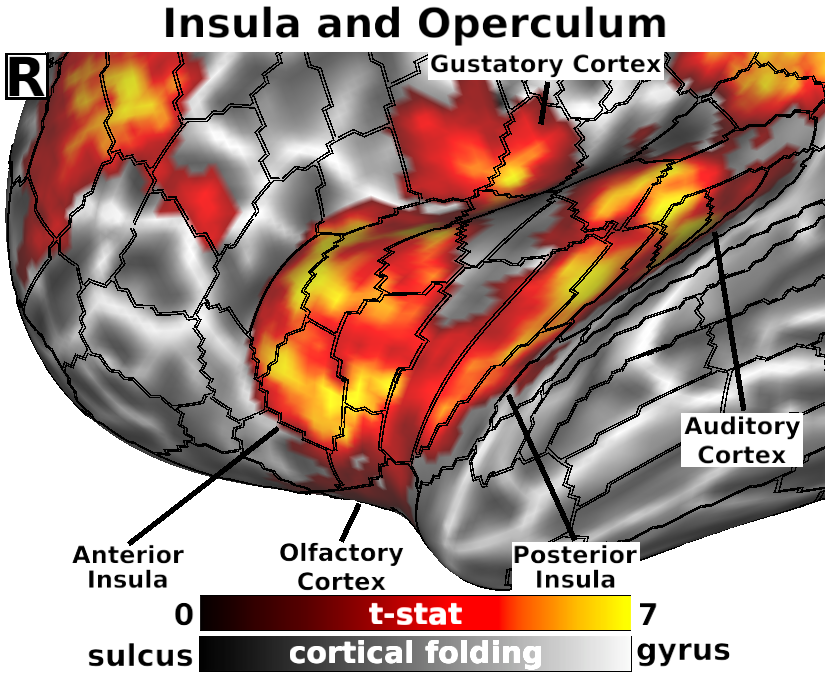
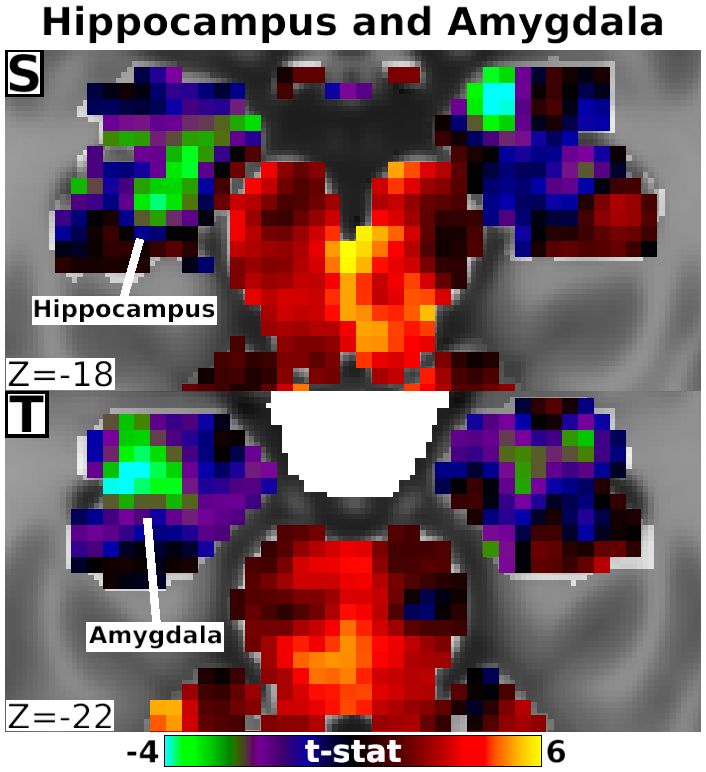
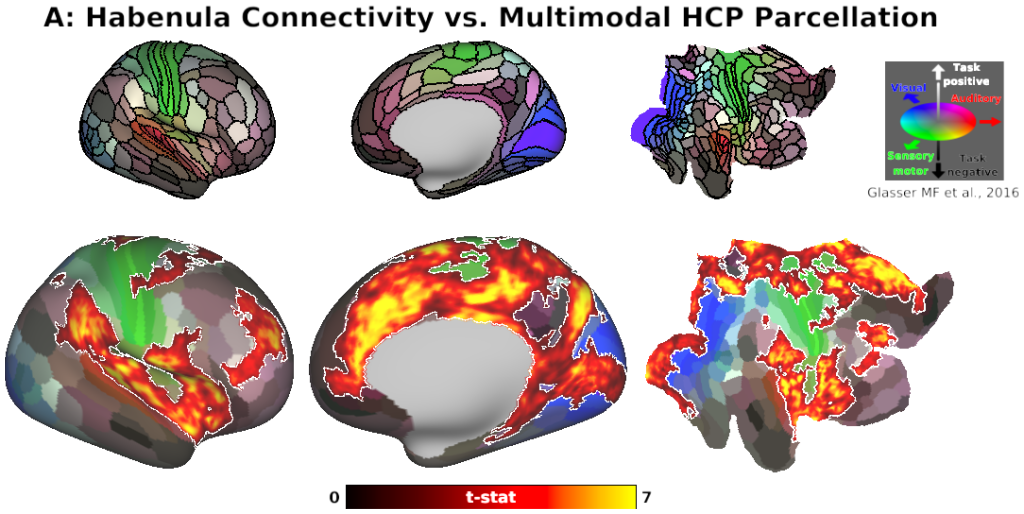
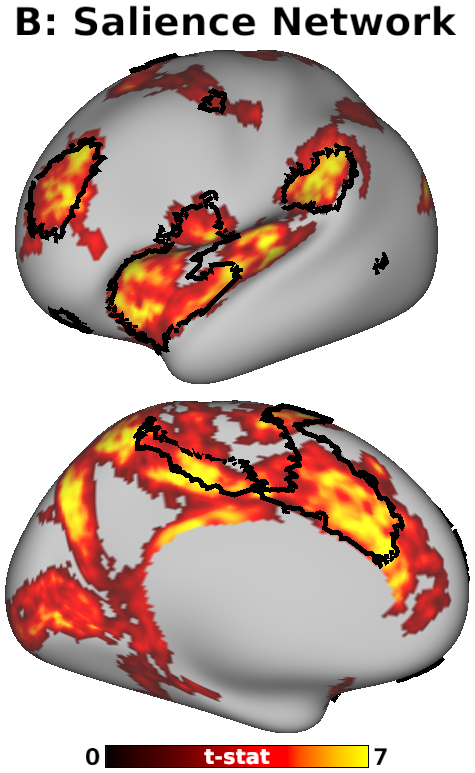
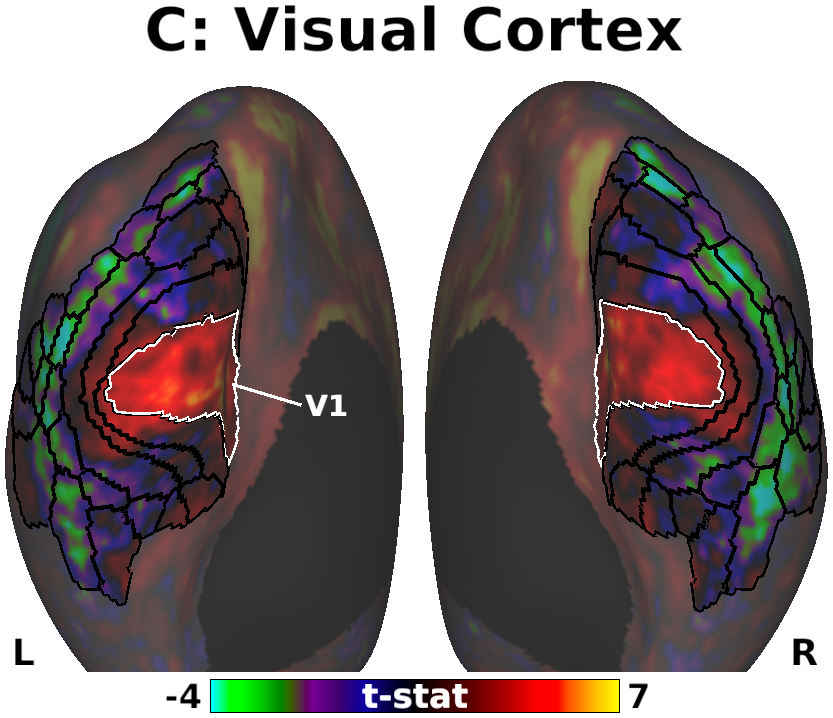
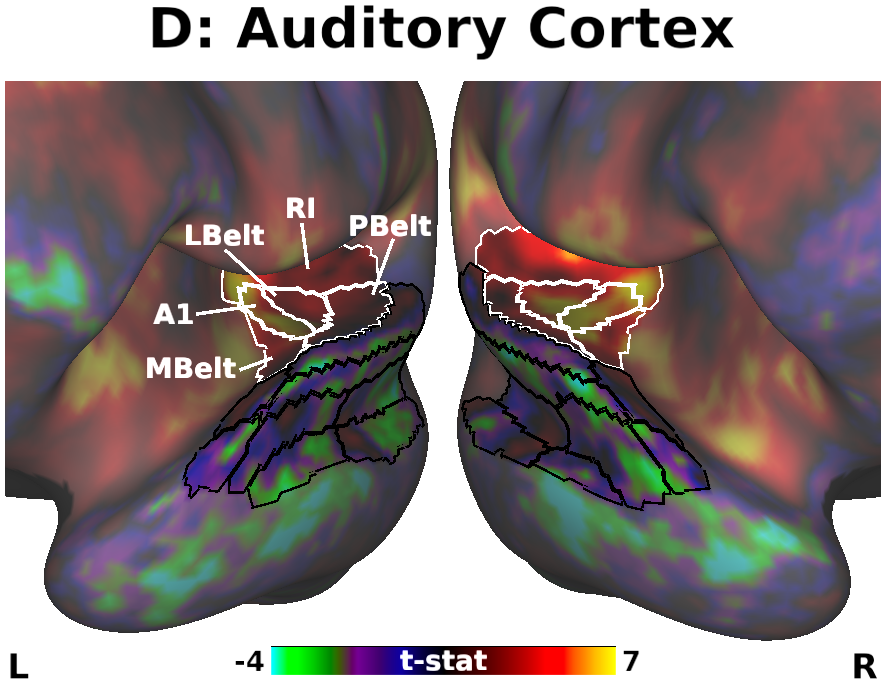
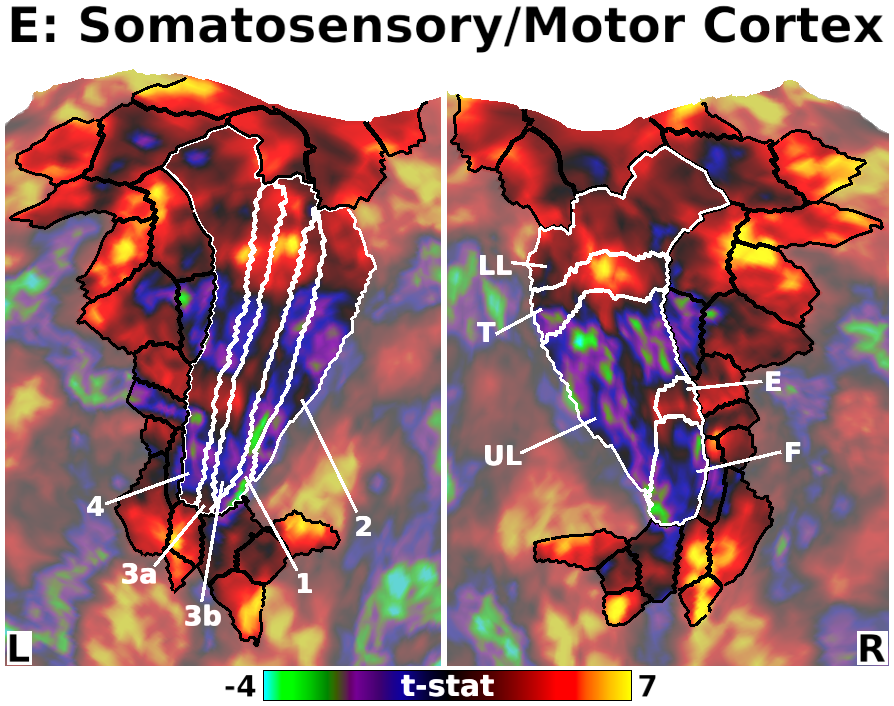
The habenula (Hb) inhibits dopaminergic reward signaling in response to negative outcomes and has been linked to numerous functional domains relevant to mental health, including reward prediction, motivation, and aversion processing. Despite its important neuroscientific and clinical implications, however, the human Hb remains poorly understood due to its small size and the associated technical hurdles to in vivo functional magnetic resonance imaging (fMRI) investigation. Using high-resolution 3T fMRI data from 68 healthy young adults acquired through the Human Connectome Project, we developed a rigorous approach for mapping the whole-brain resting-state functional connectivity of the human Hb. Our study combined an optimized strategy for defining subject-level connectivity seeds to maximize Hb BOLD sensitivity with high-quality surface-based alignment for robust functional localization and cortical sensitivity. We identified significant positive Hb connectivity with: (i) conserved brainstem targets, including the dopaminergic ventral tegmental area, serotonergic raphe nuclei, and periaqueductal gray; (ii) subcortical structures related to reward and motor function, including the nucleus accumbens, dorsal striatum, pallidum, thalamus, and cerebellum; and (iii) cortical areas associated with the Salience Network and early sensory processing, including the dorsal anterior cingulate, anterior insula, and primary visual and auditory cortices. Hb connectivity was strongly biased towards task-positive brain regions, with weak or negative connectivity observed throughout the task-negative Default Mode Network. Our study provides a detailed characterization of Hb resting-state functional connectivity in healthy young adults, demonstrating both the feasibility and clinical potential of studying the human Hb using high-resolution 3T fMRI.
PUBLICATION:
NeuroImage
- DOI:
10.1016/j.neuroimage.2019.06.015
- Benjamin A. Ely
- Emily R. Stern
- Joo-won Kim
- Vilma Gabbay
- Junqian Xu
- Icahn School of Medicine at Mount Sinai
-
Ely_Hb_connectivity.scene
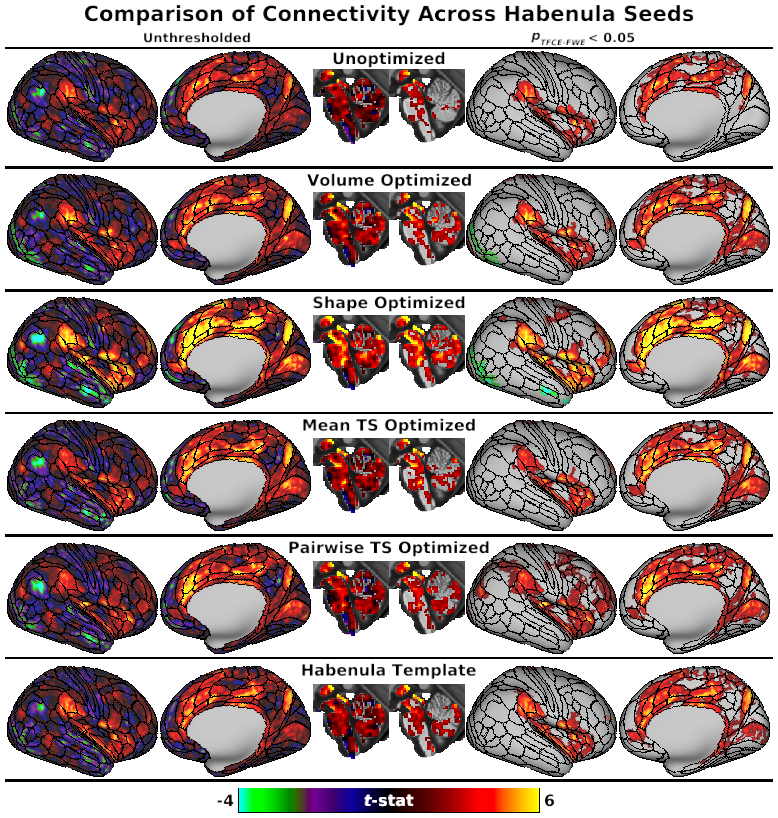
SCENES:- Fig. 3, Comparison of Connectivity Across Habenula Seeds
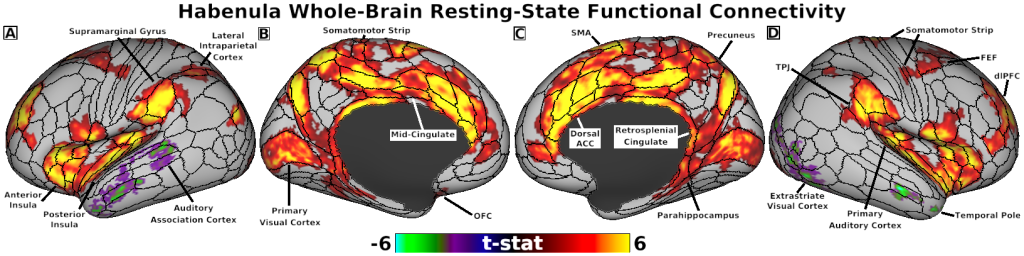
- Fig. 4, Panels A-D, Habenula Connectivity with Cortex
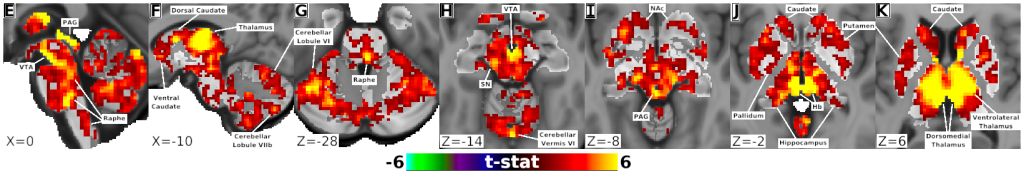
- Fig. 4, Panels E-K, Habenula Connectivity with Subcortex
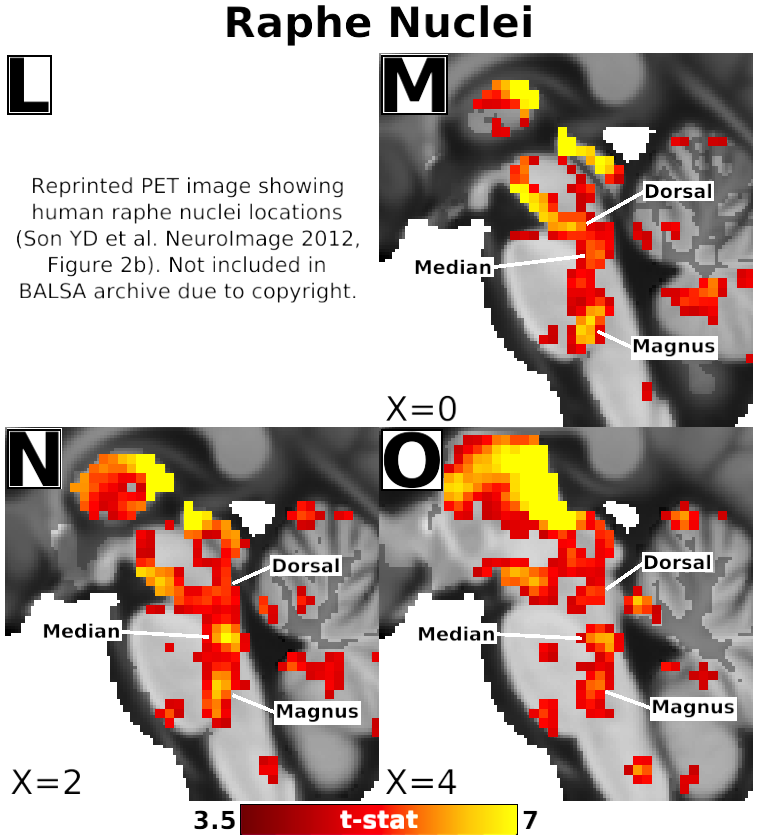
- Fig. 4, Panels L-O, Detail of Habenula Connectivity with Raphe Nuclei
- Fig. 4, Panels P-Q, Detail of Unsmoothed Habenula Connectivity with PAG and VTA
- Fig. 4, Panel R, Detail of Habenula Connectivity with Insula and Operculum
- Fig. 4, Panel S-T, Detail of Unthresholded Habenula Connectivity with Hippocampus and Amygdala
- Fig. 5, Panel A, Habenula Connectivity vs. Multimodal HCP Parcellation
- Fig. 5, Panel B, Habenula Connectivity vs. Cortical Salience Network
- Fig. 5, Panel C, Unthresholded Habenula Connectivity with Visual Cortex
- Fig. 5, Panel D, Unthresholded Habenula Connectivity with Auditory Cortex
- Fig. 5, Panel E, Unthresholded Habenula Connectivity with Somatomotor Cortex
-
Ely_Hb_connectivity_supplementary.scene
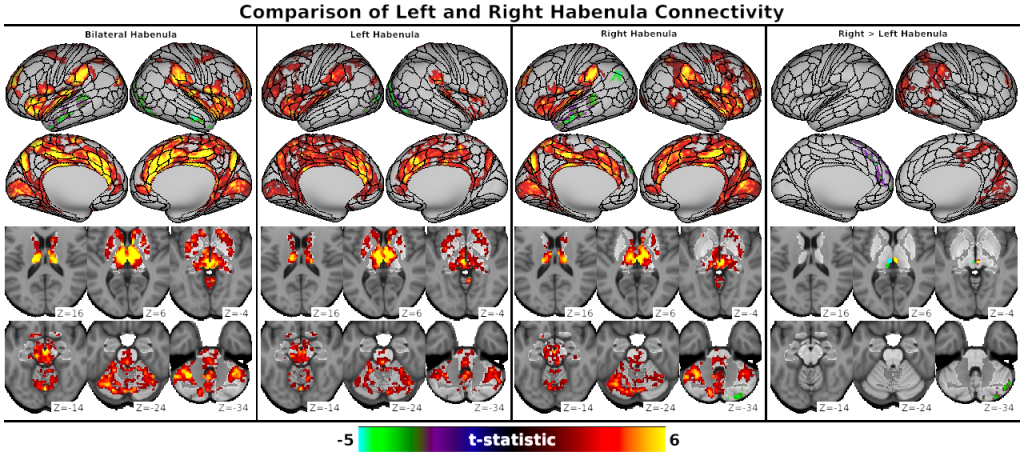
SCENES:- Fig. S2, Comparison of Left and Right Habenula Connectivity
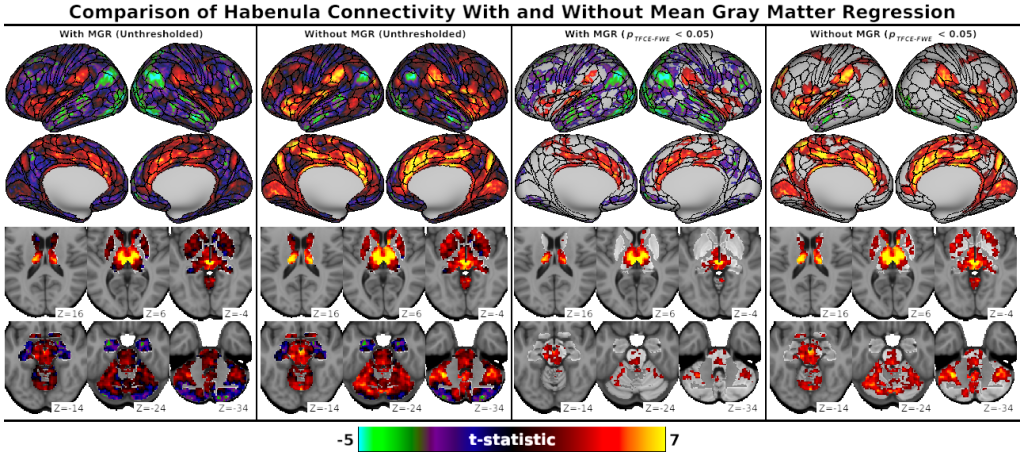
- Fig. S3, Comparison of Habenula Connectivity With and Without Mean Gray Matter Regression
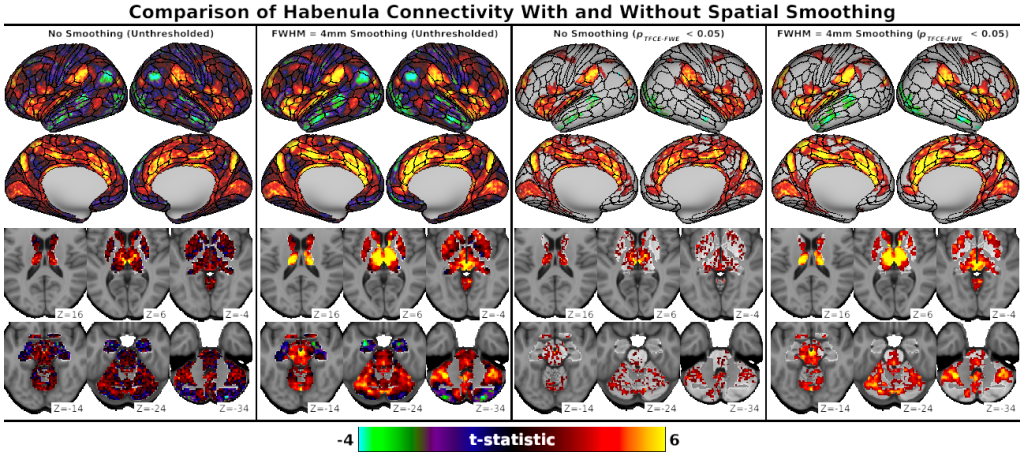
- Fig. S4, Comparison of Habenula Connectivity With and Without Spatial Smoothing
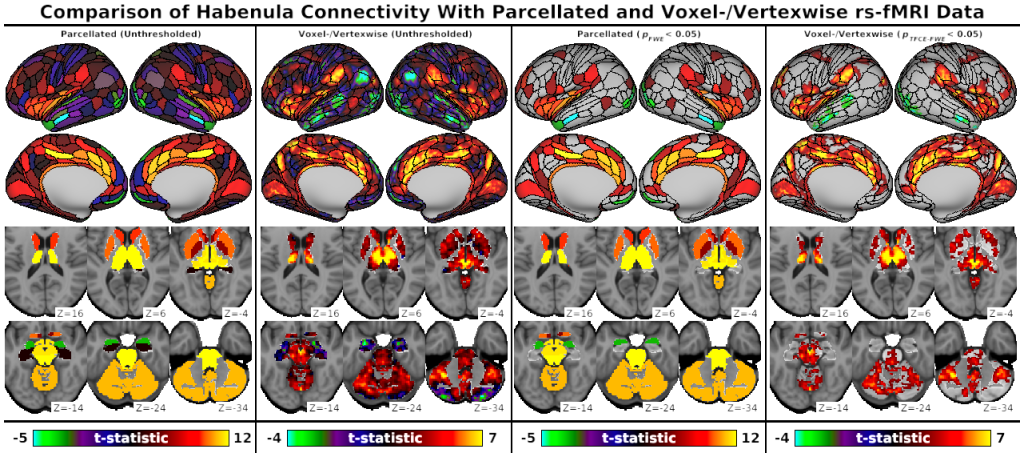
- Fig. S5, Comparison of Habenula Connectivity with Parcellated and Voxel-/Vertexwise rs-fMRI Data