study:
Temporal ICA in Functional MRI Data
SCENE FILE:
Glasser_et_al_2018_tICA_MainTextFigures
SCENE:
Figure 13
DESCRIPTION:
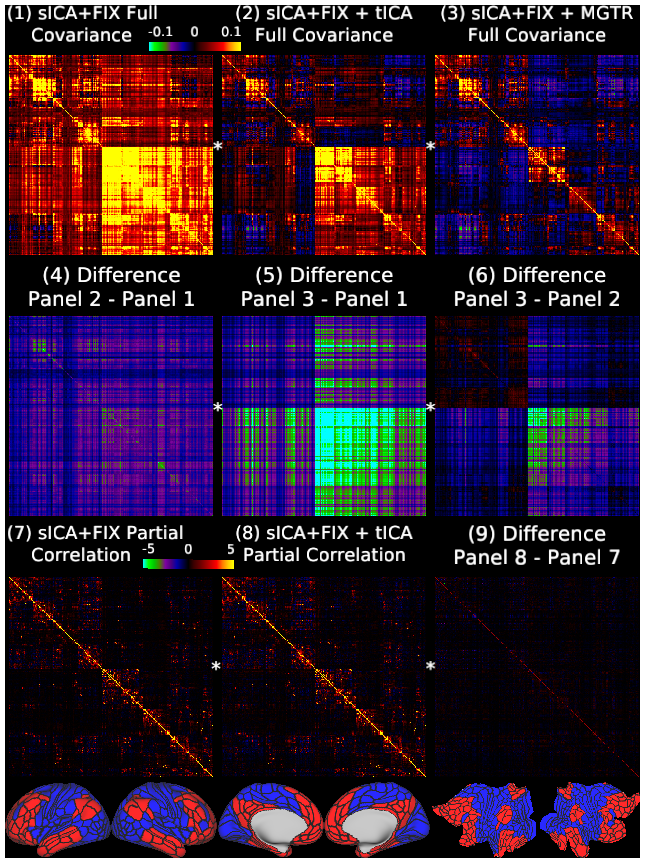
Figure 13 shows the group average full covariance matrices after sICA+FIX, sICA+FIX + tICA, and sICA+FIX + MGTR in Panels 1-3. We use covariance here because, like variances, covariances are additive and represent the absolute amount of variance shared by any two pairs of ROIs (scaled from -0.1 to 0.1 in percent BOLD for Panels 1-6). As in other figures, a global positive bias is removed by tICA cleanup (Panel 4 shows the difference between Panel 2 and Panel 1), but MGTR also removes additional signal in the bottom right quadrant of the matrix relative to the upper left quadrant with the off-diagonal quadrants in between (Panel 5 shows the difference between Panel 3 and Panel 1). Importantly, the difference between the tICA cleanup and MGTR (Panel 6 shows the difference between Panel 3 and Panel 2) is highly network specific, including small increases in cognitive/task-negative regions (bottom row, parcels shown in red) and large decreases in primarily non-cognitive/task positive regions (bottom row, blue parcels), with connections between the parcels of these two broad groups of regions showing smaller decreases. Panels 7 and 8 show that mean across subjects partial correlation regularized with ridge regression (rho=0.23, which was optimal in matching the individual matrices to the group matrix computed with no regularization; scaled Z=+/-5) is much less affected by tICA cleanup, as it already controls for global artifacts (Glasser et al., 2016b). Thus, Panel 9 (difference between Panel 8 and Panel 7) does not reveal substantial differences. The 360 cortical areas are ordered according to the same hierarchical clustering as the grey plots, and the first split, into cognitive/task negative (red) and non-cognitive and task positive (blue) regions, is shown in the bottom row and noted by a star on the netmats, with red parcels in the upper left quadrant of the netmats, and the blue parcels in the lower right quadrant. Note that it would be inappropriate to use partial correlation after MGTR, as any dataset that has zero global signal is rank deficient, because each parcel's timeseries equals the negated sum of all other parcels' timeseries.
TAGS:
Modality:T1-weighted, Modality:Myelin Map, Surface Mesh:32k fs LR, Registration:MSMAll, Modality:T2-weighted, Species:Human
