SCENE FILE:
tSNR_Prisma_vs_Connectom
SCENE:
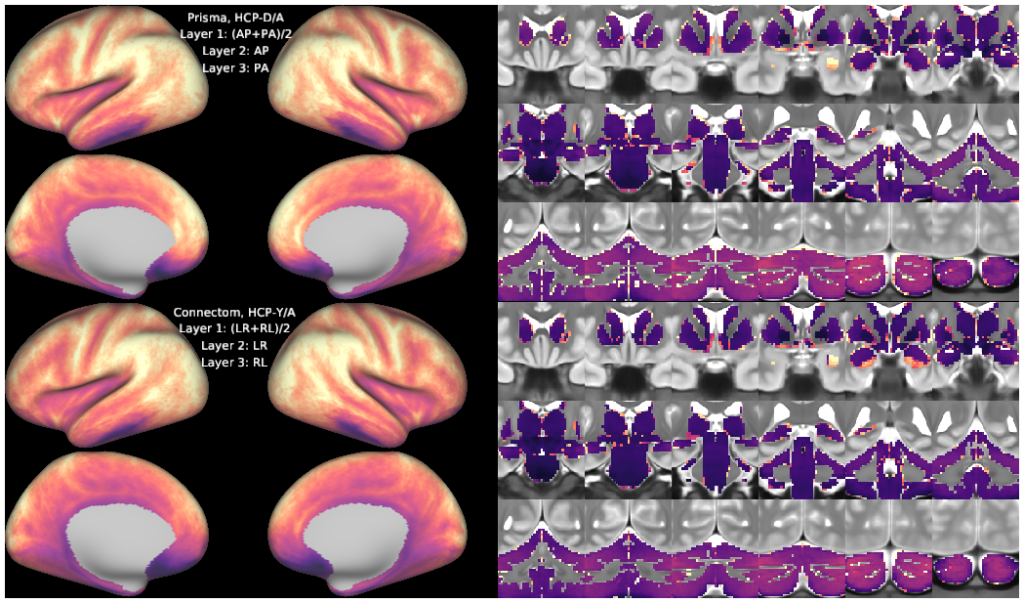
Figure 1, tSNR maps
DESCRIPTION:
This scene contains the constituent tSNR maps used to generate the density scatterplots in Figure 1 of Harms et al., 2018 (doi: 10.1016/j.neuroimage.2018.09.060).
The "HCA.tSNR.*" dscalar files are the tSNR maps from the resting-state scans collected on the Prisma scanner using the HCP-D/A protocol.
The "PCMP.tSNR.*" dscalar files are the tSNR maps from the resting-state scans collected on the same subjects on the customized 'Connectom' scanner using the HCP-YA protocol.
Maps of median tSNR (across 17 subjects, see below for further details) were computed for each separate phase-encoding polarity. The top layer shows the mean tSNR across both phase-encoding polarities that were used for a given protocol (AP/PA for HCP-D/A; LR/RL for HCP-YA).
By toggling the layers On/Off appropriately it is possible to visualize the subtle spatial effects of phase-encoding polarity on tSNR in the orbitofrontal and inferior temporal regions.
Further details (duplicated from the caption in Figure 1 of Harms et al., 2018):
tSNR maps (mean signal over time divided by the standard deviation over time) were computed for all four resting-state runs from 17 participants (age range 22-35 years; mean age=28.4; 10 females) who were scanned using both scanners/protocols (average interval: 6.4 months) after (i) processing with the HCP ‘minimal-preprocessing’ pipeline to yield a 2 mm standard grayordinate (CIFTI) representation (59412 surface vertices and 31870 ‘subcortical’ voxels, including cerebellum), (ii) truncation of the longer ‘Connectom’ rfMRI scans to a duration equivalent to the rfMRI scans collected on the Prisma (~6.4 min), and (iii) application of a Gaussian-weighted linear high-pass filter with a soft cutoff of 2000 s.
TAGS:
Modality:T1-weighted, Modality:T2-weighted, Surface Mesh:32k fs LR, Registration:MSMSulc, Species:Human
