FULL TITLE:
Cytoarchitectonic, receptor distribution and functional connectivity analyses of the macaque frontal lobe
SPECIES:
Macaque
DESCRIPTION:
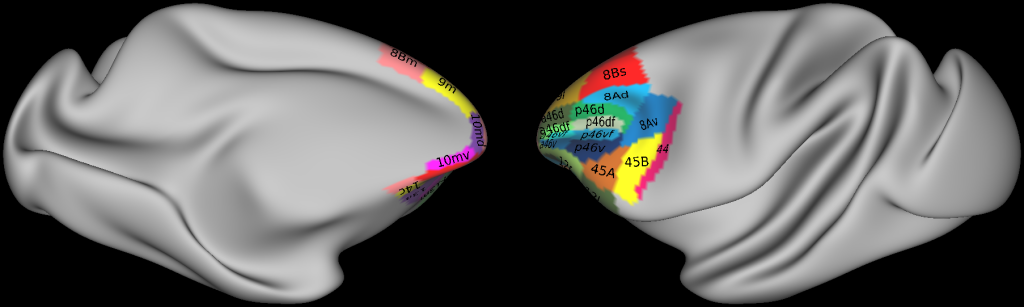
The systematic identification of 35 prefrontal areas on every 20th coronal histological section of the brain DP1, as well as silver body-stained sections of brains 11530, 11539, 11543, resulted in a map containing the location and extent of all areas. The files with the parcellation scheme are also available via EBRAINS platform of the Human Brain Project (https://search.kg.ebrains.eu/instances/Project/e39a0407-a98a-480e-9c63-4a2225ddfbe4).
ABSTRACT:
Based on quantitative cyto- and receptor architectonic analyses, we identified 35 prefrontal areas including novel subdivisions of Walker’s areas 10, 9, 8B and 46. Statistical analysis of receptor densities revealed regional differences in lateral and ventrolateral prefrontal cortex. Indeed, structural and functional organization of subdivisions encompassing areas 46 and 12 demonstrated significant differences in the interareal levels of α2 receptors. Furthermore, multivariate analysis included receptor fingerprints of previously identified 16 motor areas in the same macaque brains, and revealed five clusters encompassing frontal lobe areas. We used the MRI datasets from the non-human primate data sharing consortium PRIME-DE to perform functional connectivity analyses using the resulting frontal maps as seed regions. In general, rostrally located frontal areas were characterized by bigger fingerprints, i.e., higher receptor densities, and stronger regional interconnections. Whereas, more caudal areas had smaller fingerprints, but showed a widespread connectivity pattern with distant cortical regions. Taken together, present study provides a comprehensive insight into the molecular structure underlying the functional organization of the cortex and, thus, reconcile discrepancies between the structural and functional hierarchical organization of the primate frontal lobe. Finally, our data are publicly available via the EBRAINS and BALSA repositories for the entire scientific community.
PUBLICATION:
eLife
- DOI:
10.7554/eLife.82850
- Lucija Rapan
- Sean Froudist-Walsh
- Meiqi Niu
- Ting Xu
- Ling Zhao
- Thomas Funck
- Xiao-Jing Wang
- Katrin Amunts
- Nicola Palomero-Gallagher
- Bristol Computational Neuroscience Unit, Faculty of Engineering, University of Bristol, Bristol BS8 1UB, UK,
- C. & O. Vogt Institute for Brain Research, Heinrich-Heine-University, 40225 Düsseldorf, Germany
- Child Mind Institute
- Institute of Neuroscience and Medicine (INM-1), Research Centre Jülich, Jülich, Germany