study:
Cortical-subcortical functional networks
SCENE FILE:
ColeAnticevicNetworkPartition_MainFigures
SCENE:
Figure 6
DESCRIPTION:
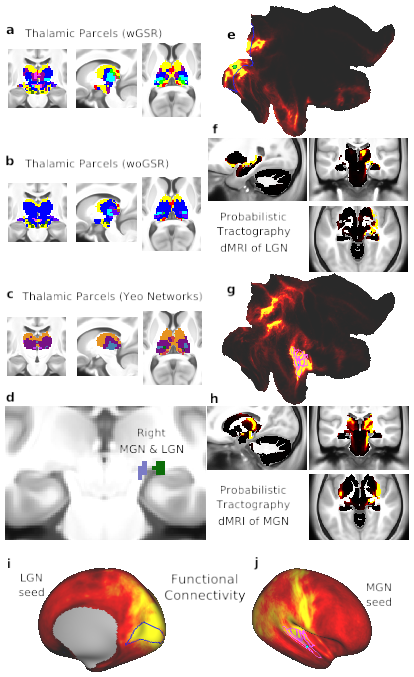
Fig. 6. Thalamic network assignment. A) Network assignment of the thalamus and ventral diencephalon from the network partition described in the manuscript. B) Network assignment of the thalamus and ventral diencephalon from the parcellation performed without GSR (woGSR). Without GSR, the auditory network assignment of the MGN was not distinguishable in the parcellation. C) Network assignment of thalamus and ventral diencephalon using cortical network parcellation from Yeo et al. (2011). Note the lack of an auditory network in the Yeo et al. (2011) partition limits the ability to map thalamus relative to the new partition reported here. D) Right LGN (green) identified using the Jülich atlas (Bürgel et al., 2006; Eickhoff et al., 2005), similar coordinates also reported in (Linzenbold et al., 2011; Marx et al., 2004; A. T. Smith et al., 2009) and right MGN (lilac) identified using the Jülich atlas. E) Probabilistic tractography (i.e. ‘structural’ connectivity) of the right primary visual cortex (V1) shown in flat cortical map. Seed grayordinate is highlighted with green dot. Cortical VIS1 network parcels are outlined in blue. Tractography results were computed from diffusion MRI data obtained from the same subjects and averaged over the entire group. F) Tractography of V1 seed to subcortex, including the right LGN. Connectivity was strongest between V1, right LGN, and other visual processing regions, including the superior colliculus and brainstem nuclei. G) Probabilistic tractography of the right primary auditory cortex, displayed in flat cortical map. Seed grayordinate is highlighted with green dot. Cortical AUD network parcels are outlined in fuchsia. H) Tractography of primary auditory seed to subcortex, including right MGN. Connectivity was strongest between right auditory cortex, right MGN, other thalamic nuclei, and auditory processing regions such as the inferior colliculi. I) Cortical functional connectivity of the bilateral LGN parcels. VIS1 parcels are outlined in blue. Right hemisphere is displayed by default; similar results are seen in the left hemisphere. J) Cortical functional connectivity of the bilateral MGN parcels. AUD parcels are outlined in fuchsia. Right hemisphere is shown by default; similar results are seen in the left hemisphere.
TAGS:
Surface Mesh:32k fs LR, Modality:T1-weighted, Species:Human
