study:
A Multi-modal Parcellation of Human Cerebral Cortex
SCENE FILE:
Glasser_et_al_2016_HCP_MMP1.0_2_SupplementaryResultsAndDiscussion
SCENE:
Figure 8
DESCRIPTION:
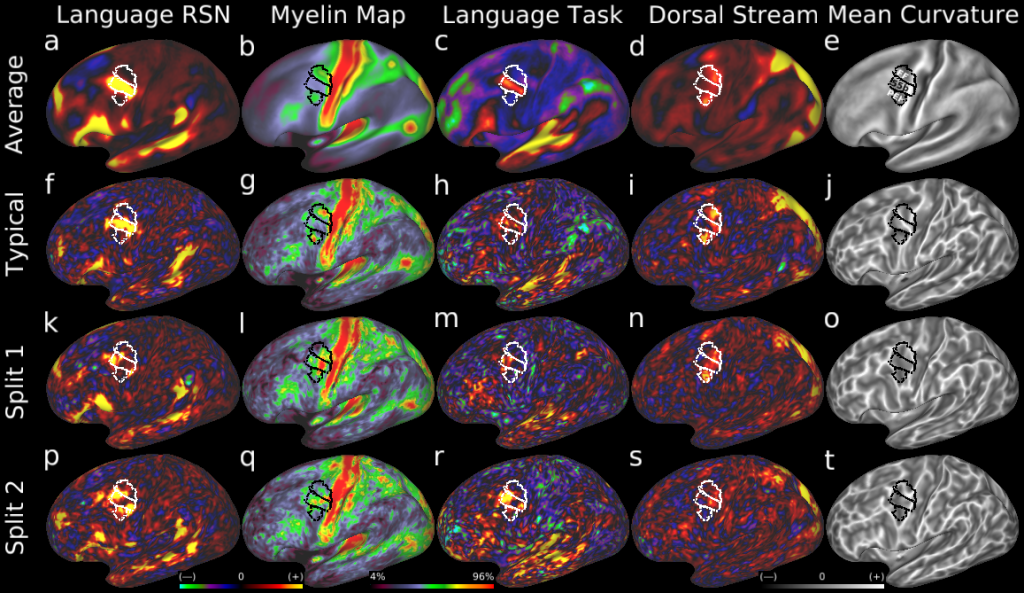
Atypical splits of area 55b. Here we show examples of a second type of topological incompatibility between the typical layout of areal features near area 55b and a minority of subjects with alternative 'split' layouts. Row 1 shows the group average areal features: a, a d=40 RSN map corresponding to parts of the language network, b, a myelin map, c, a LANGUAGE Story-Baseline task contrast beta map, d, a d=40 RSN map corresponding to the dorsal visual stream, and, e, a mean curvature map illustrating the folding pattern. The group delineation of areas 55b, FEF, and PEF are white or black lines. f-j show these maps in a typical individual subject. k-o show an atypical subject whose area 55b is split and FEF and PEF are joined. p-t show a second atypical subject with a similar pattern. In both cases the split of 55b is visible in multiple independent areal features.
TAGS:
Modality:Myelin Map, Surface Mesh:32k fs LR, Registration:MSMAll, Species:Human, Modality:T1-weighted, Modality:T2-weighted
